📥 Install | 🚀 Quick Start | ✨ Features | 📚 Documentation | 📄 Cite | 🤝 Contribute | 🔑 License
PyAerial is a Python implementation of the Aerial scalable neurosymbolic association rule miner for tabular data. It utilizes an under-complete denoising Autoencoder to learn a compact representation of tabular data, and extracts a concise set of high-quality association rules with full data coverage.
Unlike traditional exhaustive methods (e.g., Apriori, FP-Growth), Aerial addresses the rule explosion problem by learning neural representations and extracting only the most relevant patterns, making it suitable for large-scale datasets. PyAerial supports GPU acceleration (however, it is also the fastest rule miner on CPU), numerical data discretization, item constraints, and classification rule extraction and rule visualization via NiaARM library.
Learn more about the architecture, training, and rule extraction in our paper: Neurosymbolic Association Rule Mining from Tabular Data
Install PyAerial using pip:
pip install pyaerialNote: Examples in the documentation use
ucimlrepoto fetch sample datasets. Install it to run the examples:pip install ucimlrepo
Data Requirements: PyAerial works with categorical data. Numerical columns must be discretized first. This can be done using the discretization module of PyAerial. There is no need to one-hot encode your data—PyAerial handles that automatically (unlike libraries like mlxtend that require manual one-hot encoding).
PyAerial significantly outperforms traditional ARM methods in scalability while maintaining high-quality results, also on CPU:
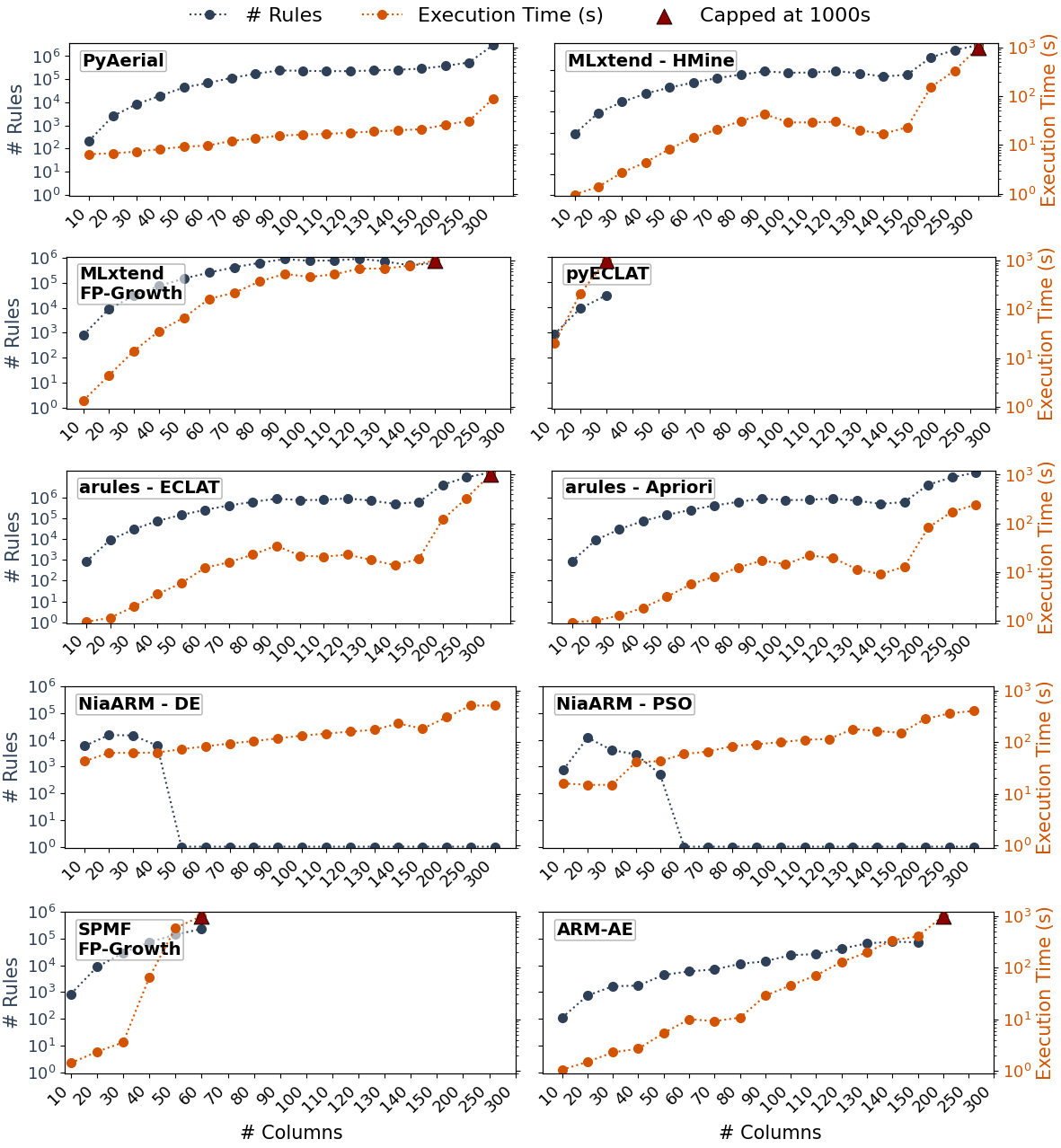
Execution time comparison across datasets of varying sizes. PyAerial scales linearly while traditional methods (e.g., Mlxtend, SPMF) exhibit exponential growth.
Key advantages:
- ⚡ 100-1000x faster on large datasets compared to standard rule mining algorithms in Python (e.g., Apriori, FP-Growth, ECLAT, ...)
- 📈 Linear scaling in training, polynomial scaling in rule extraction
- 🎯 No rule explosion - extracts concise, high-quality rules with full data coverage
- 💾 Memory efficient - neural representation avoids storing exponential candidate sets
- 🖥️ Fast on CPU - GPU is optional and only needed for very large datasets
For comprehensive benchmarking and comparisons with Mlxtend (e.g., FPGrowth, Apriori etc.), and other ARM tools, see our benchmarking paper: PyAerial: Scalable association rule mining from tabular data (SoftwareX, 2025)
The following are basic example usages of PyAerial. 📚
See full feature list | Read the complete documentation, to see the full capabilities.
from aerial import model, rule_extraction
from ucimlrepo import fetch_ucirepo
# Load a categorical tabular dataset
breast_cancer = fetch_ucirepo(id=14).data.features
# Train an autoencoder on the loaded table
trained_autoencoder = model.train(breast_cancer)
# Extract association rules with quality metrics calculated automatically
result = rule_extraction.generate_rules(trained_autoencoder, min_rule_frequency=0.1, min_rule_strength=0.8)
print(f"Overall statistics: {result['statistics']}\n")
print(f"Sample rule: {result['rules'][0]}")Output:
Overall
statistics: {
"rule_count": 15,
"average_support": 0.448,
"average_confidence": 0.881,
"average_coverage": 0.860,
"data_coverage": 0.923,
"average_zhangs_metric": 0.318
}
Sample
rule: {
"antecedents": [{"feature": "inv-nodes", "value": "0-2"}],
"consequent": {"feature": "node-caps", "value": "no"},
"support": 0.702,
"confidence": 0.943,
"zhangs_metric": 0.69,
"rule_coverage": 0.744
}Interpretation: When inv-nodes is between 0-2, there's 94.3% confidence that node-caps equals no, covering
70.2% of the dataset.
Quality metrics explained:
- Support: Frequency of the rule in the dataset (how often the pattern occurs)
- Confidence: How often the consequent is true when antecedent is true (rule reliability)
- Zhang's Metric: Correlation measure between antecedent and consequent (-1 to 1; positive values indicate positive correlation)
- Rule Coverage: Proportion of transactions containing the antecedents
- Data Coverage (in statistics): Overall proportion of the dataset covered by at least one rule
Can't get the results you're looking for?
Learn how to adjust parameters for your specific needs:
- 🎯 Parameter Tuning Guide - Quick reference for high/low support, confidence, and more
- 🔧 Troubleshooting - What to do when Aerial doesn't find rules or takes too long
- ⚙️ Advanced Tuning - Training duration and architecture optimization
📝 Note on Parameter Names: The parameters
min_rule_frequencyandmin_rule_strengthcorrespond toant_similarityandcons_similarityin the original Aerial and PyAerial papers.
Working with rules: Access rule components and metrics easily using the dictionary format:
# Example: Print all rules in a readable format
for rule in result['rules']:
antecedents_str = " AND ".join([f"{a['feature']}={a['value']}" for a in rule['antecedents']])
consequent_str = f"{rule['consequent']['feature']}={rule['consequent']['value']}"
print(
f"IF {antecedents_str} THEN {consequent_str} (support: {rule['support']:.2f}, conf: {rule['confidence']:.2f})")
# Sample output:
# IF inv-nodes=0-2 THEN node-caps=no (support: 0.70, conf: 0.94)
# IF age=30-39 AND menopause=premeno THEN breast=left (support: 0.45, conf: 0.75)
# IF tumor-size=30-34 THEN deg-malig=2 (support: 0.38, conf: 0.82)For datasets with numerical columns, use PyAerial's built-in discretization methods:
from aerial import model, rule_extraction, discretization
from ucimlrepo import fetch_ucirepo
# Load a numerical dataset (e.g., Iris)
iris = fetch_ucirepo(id=53).data.features
# Discretize numerical columns into categorical bins
iris_discretized = discretization.equal_frequency_discretization(iris, n_bins=3)
# Train and extract rules as usual
trained_autoencoder = model.train(iris_discretized, epochs=10)
result = rule_extraction.generate_rules(trained_autoencoder, min_rule_frequency=0.1)
print(
f"Found {result['statistics']['rule_count']} rules with avg support {result['statistics']['average_support']:.3f}")Example discretization output:
# Before: sepal_length = 5.1, 4.9, 7.0, ...
# After: sepal_length (ranges of values) = (4.8, 5.5], (4.8, 5.5], (6.4, 7.9], ...
Available discretization methods:
Unsupervised methods (no target variable needed):
equal_frequency_discretization- Equal-frequency (quantile) binningequal_width_discretization- Equal-width binningkmeans_discretization- K-means clustering-based binningquantile_discretization- Custom percentile-based binningcustom_bins_discretization- User-defined bin edges
Supervised methods (use target variable for classification):
entropy_based_discretization- Entropy minimization (MDLP)chimerge_discretization- Chi-square based mergingdecision_tree_discretization- Decision tree regression-based splits
Each method is documented with academic references. See the User Guide for detailed examples and references.
Focus rule mining on specific features of interest instead of exploring the entire feature space:
from aerial import model, rule_extraction
from ucimlrepo import fetch_ucirepo
breast_cancer = fetch_ucirepo(id=14).data.features
trained_autoencoder = model.train(breast_cancer)
# Define features of interest for the antecedent side, either as complete features or their specific values
features_of_interest = ["age", "tumor-size", "inv-nodes", {"menopause": 'premeno'}, "node-caps"]
# Extract rules focusing only on specified features
result = rule_extraction.generate_rules(
trained_autoencoder,
features_of_interest,
min_rule_frequency=0.1
)
print(f"Sample rule: {result['rules'][0]}")Output: Rules with specified features on the antecedent side (left side of an if-else rule):
{
'antecedents': [{'feature': 'menopause', 'value': 'premeno'}, {'feature': 'tumor-size', 'value': '30-34'}],
'consequent': {'feature': 'node-caps', 'value': 'no'},
...
}This is ideal for domain-specific exploration where you want to understand relationships involving particular features.
Learn rules with target class labels on the consequent side for interpretable classification:
import pandas as pd
from aerial import model, rule_extraction, rule_quality
from ucimlrepo import fetch_ucirepo
# Load dataset with class labels
breast_cancer = fetch_ucirepo(id=14)
labels = breast_cancer.data.targets
features = breast_cancer.data.features
# Combine features with labels
table_with_labels = pd.concat([features, labels], axis=1)
trained_autoencoder = model.train(table_with_labels)
# Generate classification rules with target class on consequent side
result = rule_extraction.generate_rules(
trained_autoencoder,
target_classes=["Class"],
min_rule_frequency=0.01,
min_rule_strength=0.7
)Output: Rules predicting class labels with quality metrics:
{
"antecedents": [{"feature": "menopause", "value": "premeno"}],
"consequent": {"feature": "Class", "value": "no-recurrence-events"},
...
}These rules can be used for interpretable inference or integrated with rule-based classifiers from imodels.
Post-filter rules to keep only high-quality ones:
from aerial import model, rule_extraction
from ucimlrepo import fetch_ucirepo
breast_cancer = fetch_ucirepo(id=14).data.features
# Train with default epochs=2 (shorter training = fewer and higher quality rules)
trained_autoencoder = model.train(breast_cancer)
# Extract rules and filter by quality thresholds
result = rule_extraction.generate_rules(
trained_autoencoder,
min_confidence=0.7, # Only keep rules with ≥70% confidence
min_support=0.1 # Only keep rules with ≥10% support
)
print(f"Found {len(result['rules'])} high-quality rules")Filtering parameters:
min_confidence: Post-filters rules to only include those with confidence ≥ this valuemin_support: Post-filters rules to only include those with support ≥ this value
PyAerial provides a comprehensive toolkit for association rule mining with advanced capabilities:
- Scalable Rule Mining - Efficiently mine association rules from large tabular datasets without rule explosion
- Automatic Quality Metrics - Rules include support, confidence, Zhang's metric, and more calculated automatically
- Smart Defaults - Short training (epochs=2) by default produces fewer, higher-quality rules
- Rule Filtering - Post-filter rules with
min_confidenceandmin_supportthresholds - Frequent Itemset Mining - Generate frequent itemsets with support values using the same neural approach
- ARM with Item Constraints - Focus rule mining on specific features of interest
- Classification Rules - Extract rules with target class labels for interpretable inference
- Numerical Data Support - 8 built-in discretization methods (unsupervised: equal-frequency, equal-width, k-means, quantile, custom bins; supervised: entropy-based, ChiMerge, decision tree)
- Customizable Architectures - Fine-tune autoencoder layers and dimensions for optimal performance
- GPU Acceleration - Leverage CUDA for faster training on large datasets
- Comprehensive Metrics - Support, confidence, lift, conviction, Zhang's metric, Yule's Q, interestingness, leverage
- Rule Visualization - Integrate with NiaARM for scatter plots and visual analysis
- Flexible Training - Adjust epochs, learning rate, batch size, and noise factors
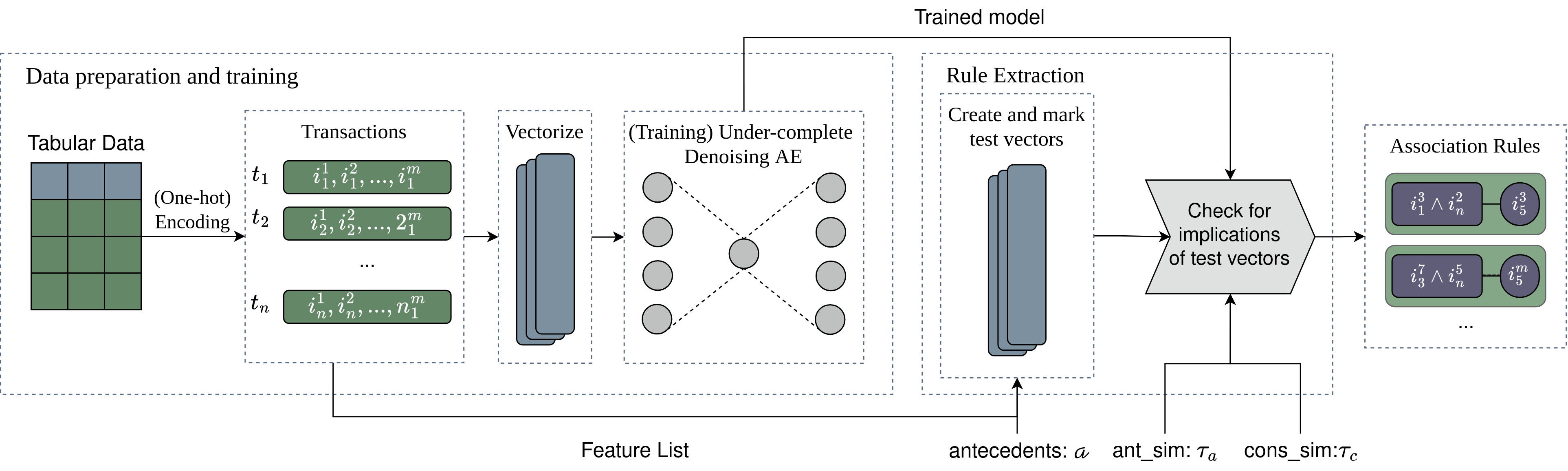
Aerial employs a three-stage neurosymbolic pipeline to extract high-quality association rules from tabular data:
Categorical data is one-hot encoded while tracking feature relationships. Numerical columns require pre-discretization ( equal-frequency or equal-width methods available). The encoded values are transformed into vector format for neural processing.
An under-complete denoising autoencoder learns a compact representation of the data:
- Architecture: Logarithmic reduction (base 16) automatically configures layers, or use custom dimensions
- Bottleneck design: The encoder compresses input to the original feature count, forcing the network to learn meaningful associations
- Denoising mechanism: Random noise during training improves robustness and generalization
Rules emerge from analyzing the trained autoencoder using test vectors:
- Test vectors are created with equal probabilities across categories
- Specific features are set to 1 (antecedents) while others remain at baseline
- Forward passes through the network produce output probabilities
- Rules are extracted when probabilities exceed similarity thresholds
- Quality metrics (support, confidence, coverage, Zhang's metric, etc.) are calculated automatically using vectorized operations
Learn more: For detailed explanations of the architecture, theoretical foundations, and experimental results, see our paper: Neurosymbolic Association Rule Mining from Tabular Data
For detailed usage examples, API reference, and advanced topics, visit our comprehensive documentation:
📚 Read the full documentation on ReadTheDocs
Documentation includes:
- Getting Started - Installation and basic usage
- User Guide - 11 detailed examples covering all features
- Parameter Tuning Guide - How to get high/low support, confidence, and control rule count
- Configuration & Troubleshooting - GPU usage, debugging, and advanced training/architecture tuning
- API Reference - Complete function and class documentation
- How Aerial Works - Understanding the neurosymbolic architecture and algorithm
If you use PyAerial in your work, please cite our research and software papers:
@InProceedings{pmlr-v284-karabulut25a,
title = {Neurosymbolic Association Rule Mining from Tabular Data},
author = {Karabulut, Erkan and Groth, Paul and Degeler, Victoria},
booktitle = {Proceedings of The 19th International Conference on Neurosymbolic Learning and Reasoning},
pages = {565--588},
year = {2025},
editor = {H. Gilpin, Leilani and Giunchiglia, Eleonora and Hitzler, Pascal and van Krieken, Emile},
volume = {284},
series = {Proceedings of Machine Learning Research},
month = {08--10 Sep},
publisher = {PMLR},
url = {https://proceedings.mlr.press/v284/karabulut25a.html}
}
@article{pyaerial,
title = {PyAerial: Scalable association rule mining from tabular data},
journal = {SoftwareX},
volume = {31},
pages = {102341},
year = {2025},
issn = {2352-7110},
doi = {https://doi.org/10.1016/j.softx.2025.102341},
author = {Erkan Karabulut and Paul Groth and Victoria Degeler},
}For questions, suggestions, or collaborations, please contact:
Erkan Karabulut 📧 e.karabulut@uva.nl 📧 erkankkarabulut@gmail.com
We welcome contributions from the community! Whether you're fixing bugs, adding new features, improving documentation, or sharing feedback, your help is appreciated.
How to contribute:
- 🐛 Report bugs - Open an issue describing the problem
- 💡 Suggest features - Share your ideas for improvements
- 📝 Improve docs - Help us make the documentation clearer
- 🔧 Submit PRs - Fork the repo and create a pull request
- 💬 Share feedback - Contact us with your experience using PyAerial
Feel free to open an issue or pull request on GitHub, or reach out directly!
All contributors to this project are recognized and appreciated! The profiles of contributors will be listed here:
Made with contrib.rocks.
This project is licensed under the MIT License - see the LICENSE file for details.