This repository provides a coupled multi-scale framework of CompuCell3D/CPM for cell-scale geometry and interactions, and a whole-cell translation kinetics model (mRNA → protein) for intracellular dynamics, with a focus on cancerology and synthetic biology applications.
- Repository contents
- Benchmarks
- Technical notes: how the coupling works
- Docker / container build
- License
- Contact
- Coupled model: CPM (shape, contacts, motion, division) ↔ whole-cell translation kinetics (ribosomes, tRNAs, protein production)
- Custom CC3D plugins (C++): geometry and mechanics (including a Maxwell viscoelastic medium)
- Benchmarks designed to demonstrate scientific relevance and computational scalability
- Container workflow based on a pre-built DockerHub image:
gdavid57/cc3d-compile
Benchmark folder: benchmarks/01_spheroid_scalability/
Scales the coupled model from 1 → 1024 cells to quantify runtime and assess feasibility for tumor spheroid simulations (hundreds to thousands of cells) on multi-core machines. The benchmark records timing vs. cell count as the spheroid grows.
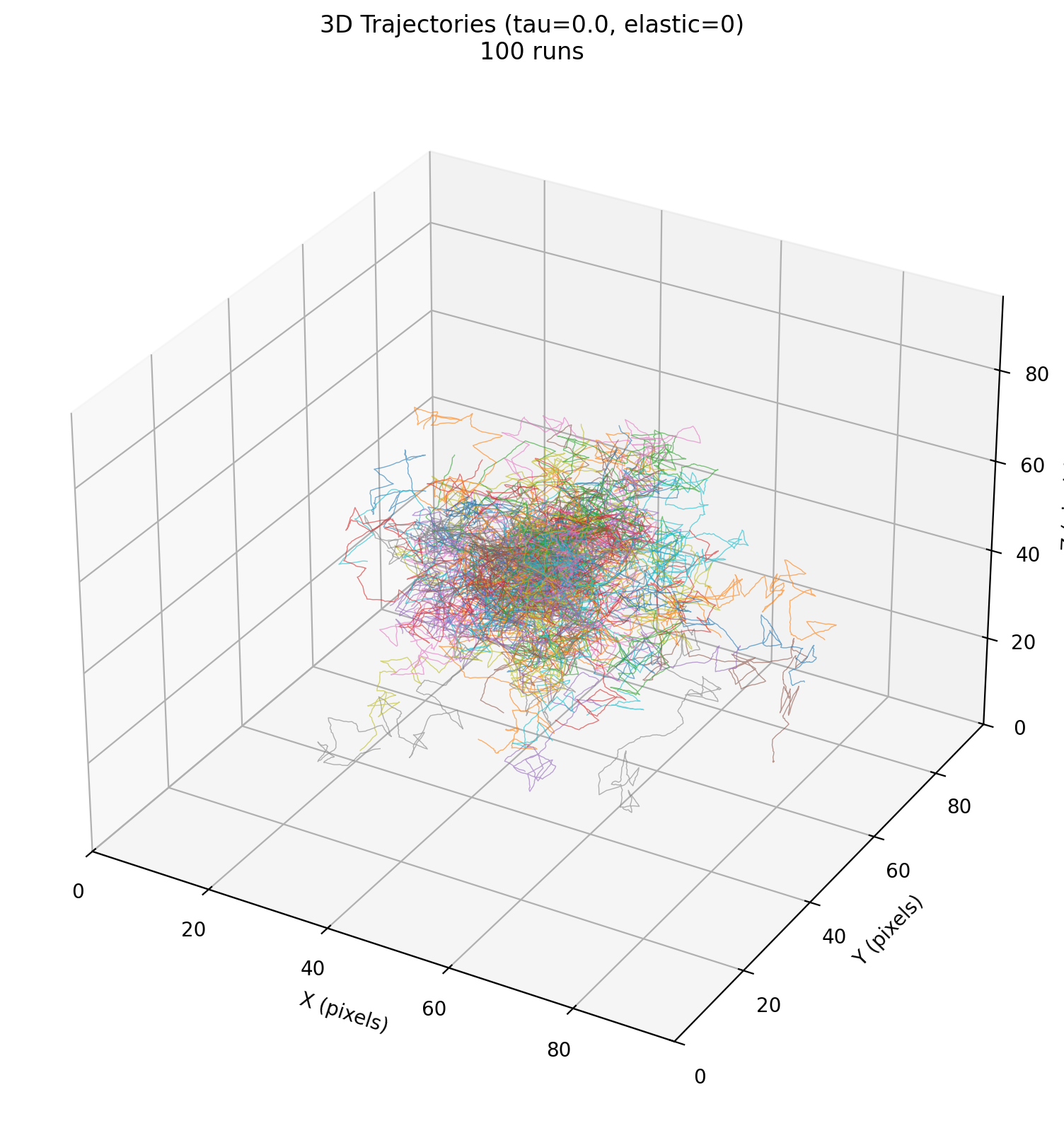
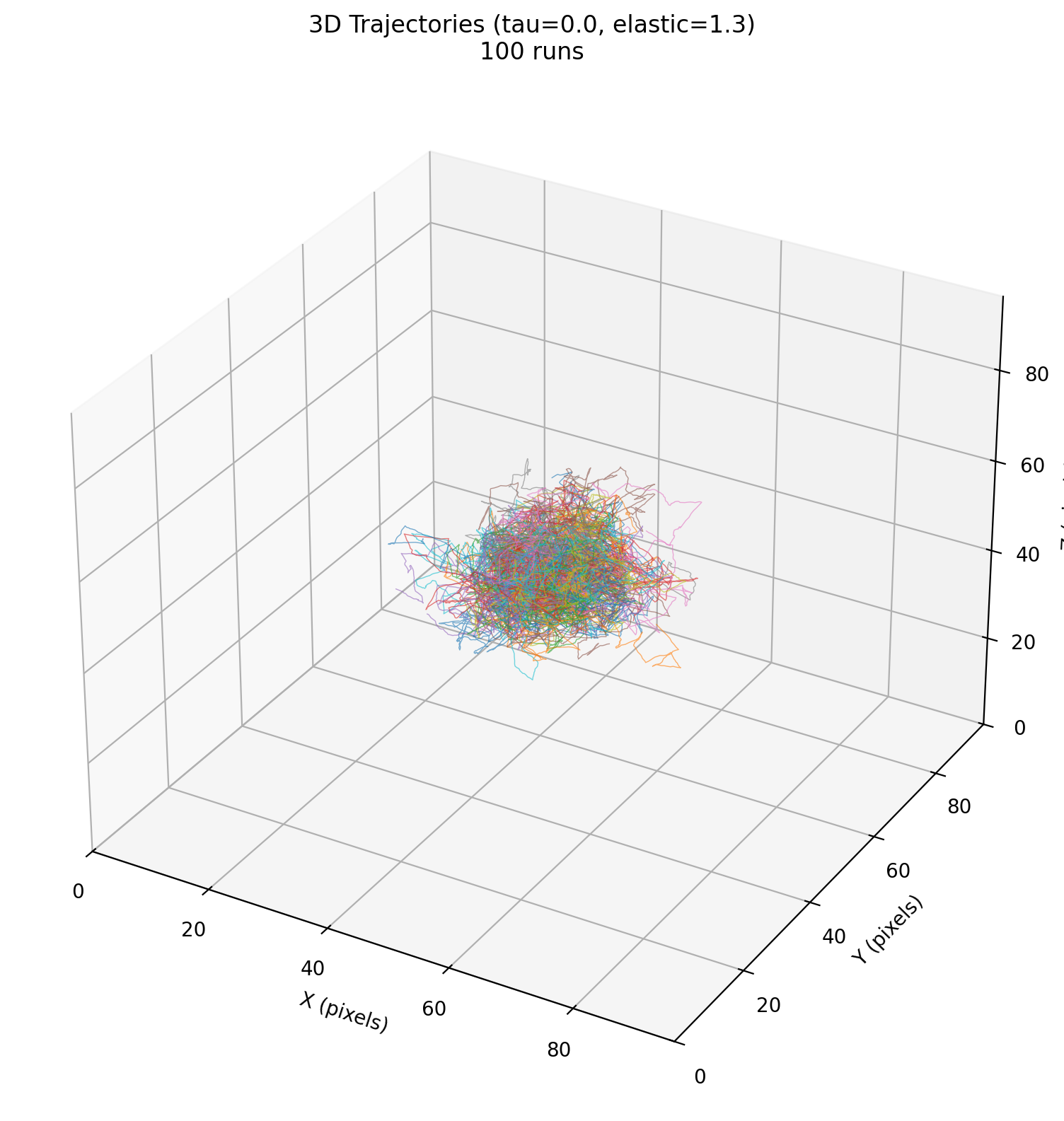
Benchmark folder: benchmarks/02_mechanical_confinement/
Studies the statistics of a single cell’s constrained motion under a Maxwell viscoelastic medium (elastic response + stress relaxation), as a proxy for tumor cell confinement in surrounding tissue. The benchmark runs multiple replicates and analyzes trajectories/MSD across parameter sweeps (elasticity, relaxation time).
Benchmark folder: benchmarks/03_bacteria_whole_cell/
Couples a rod-shaped bacterium geometry (via CC3D plugins) with the intracellular translation kinetics model to track growth and division under different parameterizations. This benchmark is a stepping stone toward Engineered Living Materials (ELMs) applications where intracellular state impacts collective behavior.
The commands below use the pre-built DockerHub image gdavid57/cc3d-compile and mount your local checkout into the container (outputs are written on your machine).
# 0) One-time setup
docker pull gdavid57/cc3d-compile
# 1) Benchmark 01 — Tumor spheroid scalability
docker run --rm -it \
-v "$(pwd)":/work \
-w /work/benchmarks/01_spheroid_scalability \
gdavid57/cc3d-compile \
conda run -n cc3d_compile python multiprocessing_wc_script.py default_config.json
# 2) Benchmark 02 — Maxwell confinement: full sweep
docker run --rm -it \
-v "$(pwd)":/work \
-w /work/benchmarks/02_mechanical_confinement \
gdavid57/cc3d-compile \
conda run -n cc3d_compile python cell_brownian_motion_maxwell_constraint.py --sweep
# 3) Benchmark 03 — Bacteria whole-cell
docker run --rm -it \
-v "$(pwd)":/work \
-w /work/benchmarks/03_bacteria_whole_cell \
gdavid57/cc3d-compile \
conda run -n cc3d_compile python automated_wc_cc3d_script.py default_config.json
# 4) Generate all figures (after running benchmarks)
docker run --rm -it \
-v "$(pwd)":/work \
-w /work \
gdavid57/cc3d-compile \
conda run -n cc3d_compile python scripts/generate_all_figures.pyAt a high level, the coupling alternates between:
- CPM/CompuCell3D step(s) to update cell geometry and interactions on the lattice
- Whole-cell translation kinetics to update intracellular state as a function of cell state (e.g., volume), producing outputs such as protein counts
To keep the coupled simulation tractable as the number of cells grows, the benchmarks use multiprocessing for the intracellular component: each cell’s translation step can be computed independently for a given Monte Carlo step (MCS), so per-cell updates are dispatched in parallel to worker processes and the updated state is then re-attached to the corresponding CC3D cell.
For image build details (from source), contexts and options, see:
Proprietary — All rights reserved. See LICENSE.
Dr Adélaïde Raguin
adelaide.raguin@u-bordeaux.fr